PetaSuite
PetaGene’s compression software addresses challenges caused by growing volumes of genomics data. It achieves savings of between 60% and 90% in both storage costs and data transfer times compared to BAM and gzipped FASTQ files - this is a 96% reduction compared to raw FASTQ files. It transparently integrates with existing storage infrastructure and bioinformatics pipelines. PetaSuite is a set of scalable complementary software tools that significantly reduce the size and cost of NGS data for storage and transfer.
PetaLink Cloud Edition
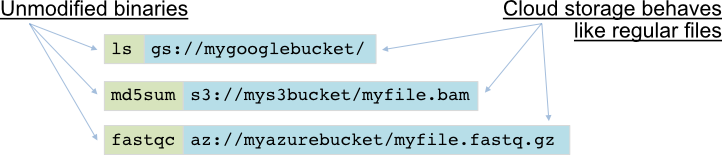
PetaLink Cloud Edition works with both uncompressed and compressed data. It enables a user’s software tools and pipelines to seamlessly integrate with a wide variety of cloud platforms without modification. AWS, Azure, GCP, private cloud and hybrid cloud are all supported transparently. PetaLink allows data to be compressed and streamed to object storage in a single step. It also provides streaming random access to data in object storage as if it were regular files, and does on-the-fly decompression if necessary. This avoids the need to download data from object storage before use.
PetaSuite Protect
PetaSuite Protect is an encryption, access management and auditing tool for genomic data. When sharing files with internal or external collaborators, you can control fine-grain access, demonstrate compliance and enable deep-auditing of usage for regular genomic data files.
The graphical user interface makes it easy to encrypt files using FIPS 140-2 compliant AES-256 regional encryption and manage user access.
Features
Lossless Compression
Our robust, high performance FASTQ.gz and BAM compression will decompress back to exactly match the original file content. There is full validation and MD5 matching, meaning that not only is the internal content of FASTQ.gz and BAM files preserved, but the gzip wrappers will exactly match, allowing simpler archiving procedures to be used.

PetaLink
PetaLink is a powerful virtual file access system. It enables migration of BAM and FASTQ.gz data to more efficient compression formats. For example, after the PetaSuite binary has been used to losslessly compress a BAM file, validate that all data in the BAM has been preserved, and remove the original BAM file, PetaLink makes available a high performance virtual BAM file view of the compressed file, with the filename of the original file, in the same location. This virtual file can then be used just like the original BAM file by Linux toolchains, pipelines and genome browsers transparently.
The Cloud Edition of PetaLink also allows files stored remotely in the cloud to be accessed as if they are local, without downloading them first!
Bayescal Quality Score Refinement
BayesCal uses a Bayesian approach to calculate a more complete posterior estimation of sequencer error. Genotyping accuracy is preserved across the ROC curve, with a net increase. Improved compression is a side effect, increasing compression ratios by a further 30-70% compared with straight lossless compression.
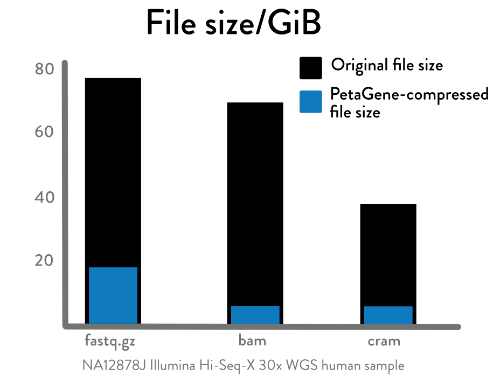
PetaGene lossless compression ratios, compared with CRAM
| Source data (human 30x WGS) |
Pipeline | PetaGene % savings |
CRAM (latest) % savings |
|---|---|---|---|
| FASTQ.gz, HiSeq X | 67% | Not applicable | |
| FASTQ.gz, NovaSeq | 77% | Not applicable | |
| BAM, HiSeq X | BWA-mem only | 55% | 47% |
| BAM, HiSeq X | GATK | 81% | 33% |
| BAM, NovaSeq | Isaac only | 64% | 57% |
| BAM, NovaSeq | BWA-mem only | 69% | 58% |
| BAM, NovaSeq | GATK | 91% | 33% |
Note: using PetaGene’s optional BayesCal quality score refinement increases the compression ratio by a further 30%–70%.